I have had little success with DSF and FP assay development for USP5 Zf-UBD to identify small molecule ligands of this domain. I decided to try 19F NMR as a potential screening assay. NMR spectroscopy can be used to visualize changes in protein spectra upon addition of low complexity molecules as well as to quantify binding affinity between the protein and compounds1. USP5 Zf-UBD is well suited for 19F NMR due to its small size as well as the presence of two tryptophan residues, one of which is in the binding site and is involved in co-ordinating ubiquitin binding. The tryptophan residues are good targets for a fluorine label. Before screening, I first had to express and purify the fluorinated protein. Experimental details are on Zenodo.
The aim was to purify protein with 5-fluoro-tryptophan (5F-W) incorporated at both sites in the protein molecule. I was able to purify almost 13 mg of protein from a 1 L test expression. Label incorporation at both tryptophan residues was verified by mass spectrometry. I also checked that the fluorinated protein was folded and bound the native ligand using DSF and FP assays.
Before 19F NMR compound screening, I had to check if there are perturbations in at least one of the 19F resonances using a ubiquitin peptide that binds to the USP5 Zf-UBD. I saw two well-resolved peaks of equal intensity for the protein. The addition of the ubiquitin peptide resulted in a significant chemical shift and decrease in peak area for the peak at ~118 ppm (Figure 1). This is exciting news-I can detect ligand binding! Next, I will use 19F NMR to screen compounds against USP5 Zf-UBD. More on this later!
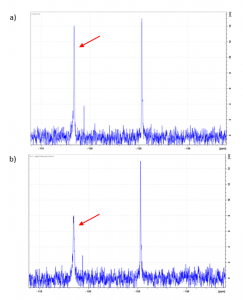
Figure 1. 19F NMR spectra of a) 50 µM 5F-W USP5171-290 and b) 1:3 5F-W USP5171-290: LRLRGG
1Gee, T. C., Arntson, E. K., Urick, K. A., Mishra, K. N., Hawk, M. L., Wisniewski, J. A., & Pomerantz, C. W. (2016). Protein-observed 19F-NMR for fragment screening, affinity quantification and druggability assessment. Nature Protocols. 11(8): 1414-1427

One Reply to “5-fluoro-tryptophan USP5 Zf-UBD Growth & Purification for 19F NMR Screening”