The experimental set-up and results covered in this post can be found here: doi.org/10.5281/zenodo.1420537
I was previously using an HTT overexpression model to look at any changes in proteins related to HTT or its function. Unfortunately, none of the related proteins I looked at showed any significant change in protein levels with HTT overexpression. There are several reasons for this: the HTT overexpression might not be strong enough (especially for the larger constructs of D4 and D6, which are harder to express) or the level of HTT protein in the cells is already saturated, hence an overexpression does not further affect these proteins of interest. I therefore decided to use a knockdown model instead, which might provide more insight into the functions and effects of HTT.
A slightly different system was used to generate the HTT knockdown models, compared to the HTT overexpression model. For the overexpression model, a baculovirus system was used. In the knockdown model, a lentivirus system was used.
Lentiviruses can be easily produced through the transfection of HEK293T cells with several plasmids which code for different components of the lentivirus. The HEK293T cells, which are very easy to transfect, can then produce the various components of the lentivirus and assemble the lentiviruses, releasing them into the cell media. As HEK293T cells are grown adherently, these lentiviruses can be harvested by simply collecting the supernatant. When the plasmids encoding the different components of the lentivirus are transfected together with a plasmid encoding an shRNA for the gene of interest (in this case, HTT), the lentiviruses produced have the ability to disrupt the gene of interest through RNA interference (RNAi) when they transduce the target cells.
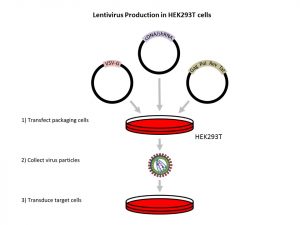
Figure adapted from Figure 4 of Nucleic Acid Delivery: Lentiviral and Retroviral Vectors by Erin P O’Keefe; Princeton University, United States (MATER METHODS 2013;3:174; dx.doi.org/10.13070/mm.en.3.174).
Upon transduction of the target cells, the lentivirus delivers the plasmid encoding the shRNA to the nucleus. The plasmid produces the shRNA, which is processed by the target cell, leading to the production of a sequence complementary to that of the mRNA of the gene of interest, HTT. The binding of this complementary sequence to the mRNA of the gene of interest causes the formation of the RNA-induced silencing complex (RISC). The RISC, consisting of both the complementary sequence from the shRNA and the mRNA of the gene of interest, is then degraded, preventing the expression of the target protein, which is HTT in this case, leading to the knockdown of HTT. As the plasmid in the target cell continuously produces the shRNA, the mRNA is continuously degraded and the cells are stably knocked down for HTT.
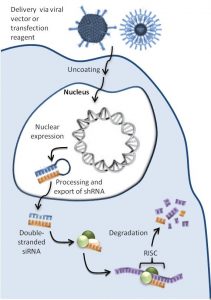
Figure from Figure 1 of DNA-directed RNA interference (http://www.wikiwand.com/en/DNA-directed_RNA_interference).
Using this method, I managed to establish 3 stable HTT knockdowns using 3 independent shRNA constructs in HeLa cells.
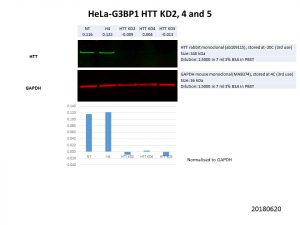
As you can see, the levels of HTT are much lower in the HTT knockdown cells compared to the no treatment and H4 (luciferase) controls. I will next use these cell lines to identify any changes in expression of HTT-related proteins.

HTT staff, Think crazy. Have you ever wondered if microplastics (< 5 mm) are able to cross the blood-brain barrier? People have been ingesting microplastics since the mass-production of plastic bottles and their commercial usage beginning in the 1950's. Side note: People that drink wine, drink wine from glass bottles; drinking wine is part of the Mediterranean diet (fresh vegetables/no plastics (bisphenol A) = less neurodegeneration). Are the chemical compounds associated with microplastics (polypropylene and polyethylene/organophosphorus esters/phthlates) found in the brain tissue/cells of people with neurodegenerative/carcinogenic brain disorders? Gosh, fish ingest microplastics, people eat fish containing microplastics. Enjoy 🙂