It’s been a very busy few weeks, a fair amount of lab work but also a fair bit of outreach work (which is the other part of my job) and so I’ve sadly neglected to write my poor blog post. However, now I’ve got the chance to write a bit about what I’ve been up to.
Previously I’d talked about how using a pulldown it looked like ACVR2 was being pulled down in complex with XIAP. To explore this further I attempted to purify the complex of ACVR2 with XIAP using protein I had already purified.
First I had to remove the GST tag from XIAP which I did by incubating with Tev protease overnight. Then I had to purify the XIAP from the GST tag and the uncleaved protein.
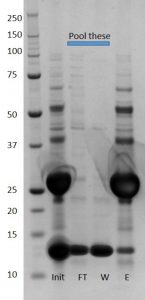
A mix of cleaved and uncleaved can be seen in the initial sample (init) and in the final elution (E) while only a strong lower band for XIAP alone is seen in flow through (FT) and wash (W)
I mixed ACVR2 with excess XIAP and ran it down a size exclusion column. However it appears that the two proteins do not have a high enough affinity to be co-purified in this fashion.
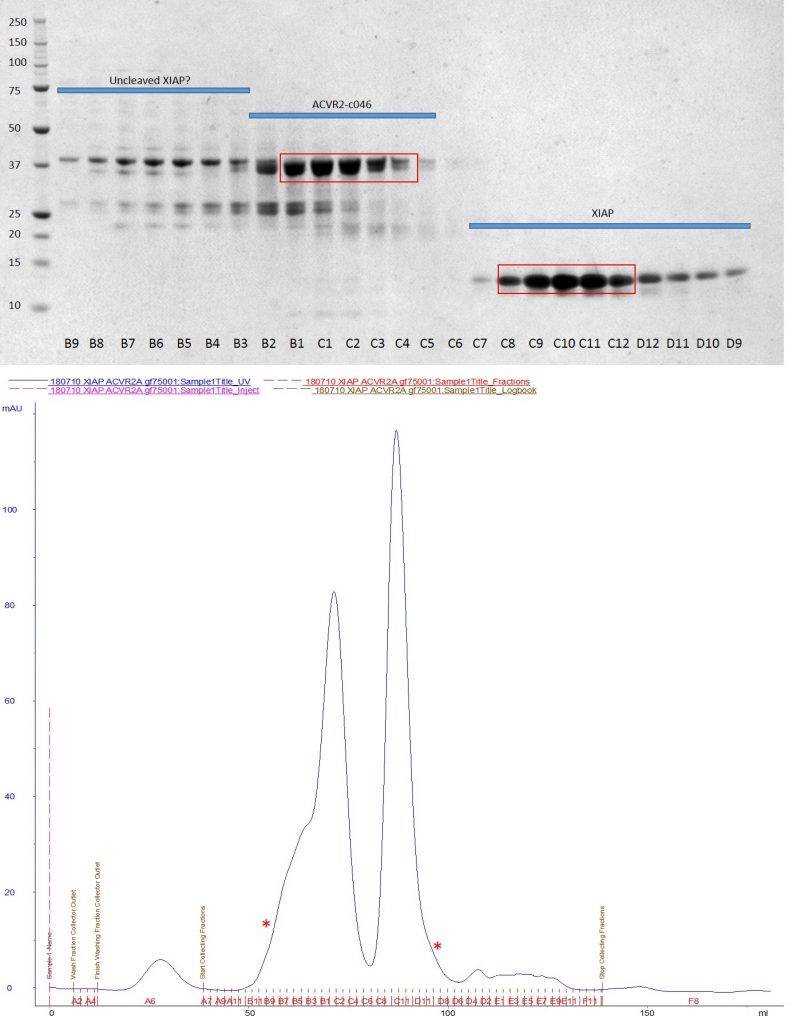
Top: SDS-PAGE of GF75 size exclusion run showing no complex formation between XIAP and ACVR2. Bottom: Corresponding UV trace from the AKTA
Instead I mixed the proteins at a 1:1 ratio, concentrated them down, added 0.5mM LDN-193189 and set up crystal plates using four coarse screens at both 4°C and 20°C.
I found two crystals which I mounted but neither of them were very promising. I sent them on the next available trip to the Diamond light source and neither of them diffracted.
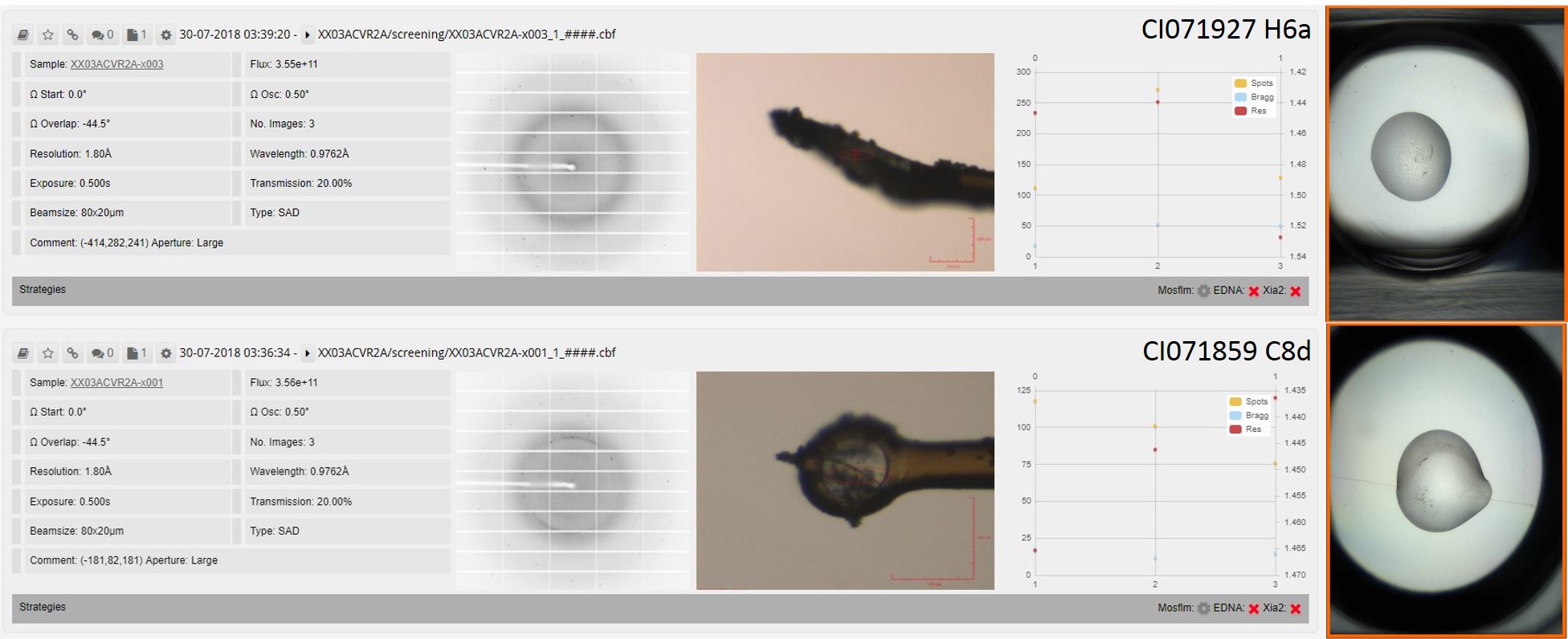

Crystal drops, mounted crystals and diamond screening data for ACVR2/XIAP complex.
So… back to the drawing board.
The next step is to try some different constructs of XIAP as the one I used was just one domain. I’ve obtained a plasmid for a full length XIAP-GST tagged construct from a collaborating lab which I have transformed into both MACH1 and BL21-DE3 cells and expressed 6l in LB from the BL21-DE3 stock ready for purification. The collaborator also has other constructs of XIAP with different domains that we can try after trying the full length to try and narrow down which domain (if any) interact with the Type I and Type II protein.
More details on all of this can be found over at Zenodo.

Perhaps the proteins don’t interact and you’re wasting your time.
This is very possible so I’m not going to waste more time with this construct of XIAP which contains only the BIR2 domain of the protein (but was the most readily available to test at the time within the SGC since we had the clones for it already). The other construct of XIAP that I’ve now got from our collaborator is of the full length protein which consists of three BIR domains and a RING domain. Given this protein is expressed in E.coli and so is fairly quick and easy to produce it seems worth repeating the pull-down before moving on completely. I’ve still got protein left from the last pull down I did so the only protein I need to purify is the full length XIAP. Hopefully this won’t take too long!