These days CRISPR gene editing needs little introduction. From humble origins as a bacterial defence system evolved to thwart viral invaders, CRISPR-based technologies have recently made headlines for their promise to revolutionize medicine. While there is a lot of excitement surrounding the use of gene editing in the clinic, it is also an incredibly valuable tool for studying the function of proteins in cells. For my project, I will be using CRISPR to make genetic changes to the NSD3 gene and study the outcome, beginning first by using the technology to create NSD3 knock out cell lines.
There is an ever-expanding catalogue of CRISPR reagents available to scientists. I’ve decided to use a method that involves the in vitro (in a tube) assembly of ribonucleoproteins (in this case Cas9 + annealed tracrRNA:crRNA – RNPs), which can then be introduced into a cell via lipofection or electroporation. Using RNPs as opposed to viral delivery or plasmid-based transfections is advantageous as the expression of Cas9 is transient limiting off-target effects and it is compatible with many different cell types.
Using this approach, the first step would be to either buy or purify recombinant Cas9. Since I work at the SGC, where expertise in protein production is abundant, I opted to purify recombinant Cas9 using Jennifer Doudna’s expression plasmid from AddGene (# 69090). With help from Rachel Harding and Mani Ravichandran, I was able to make highly pure, low-endotoxin, active Cas9 protein for my future editing experiments (see extended details of the protein prep & QC on zenodo). Next steps – getting the RNPs into cells, testing for INDELs at the cut site, and screening clones for NSD3 disruption.
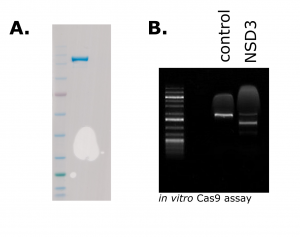
Figure 1. Purified Cas9 (A). 1 µg of purified Cas9 run on an SDS-PAGE gel and stained with Coomassie Blue (B) In vitro activity assay showing specific cleavage of NSD3-short PCR-amplified DNA when the Cas9-RNP is assembled with NSD3-targeting crRNA, but not the non-targeting control RNA.
Acknowledgements: A big thank you to Rachel Harding and Mani Ravichandran at the SGC Toronto for help optimizing the protocol as well as running instrumentation for size exclusion and cation exchange. I would also like to thank Dr. Jennifer Doudna for sharing her Cas9 expression construct on addgene (Addgene plasmid # 69090) [1].
References:
1. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. Lin S, Staahl BT, Alla RK, Doudna JA. Elife. 2014 Dec 15;3:e04766. doi: 10.7554/eLife.04766. 10.7554/eLife.04766 PubMed 25497837

I’m interested in your research, and I I wonder if you have tested the purity of cas 9. More precisely, have you tested the purity of cas 9 with a reversed-phase column?